Wei Chen
Professor of Pediatrics
University of Pittsburgh
Principal Investigator
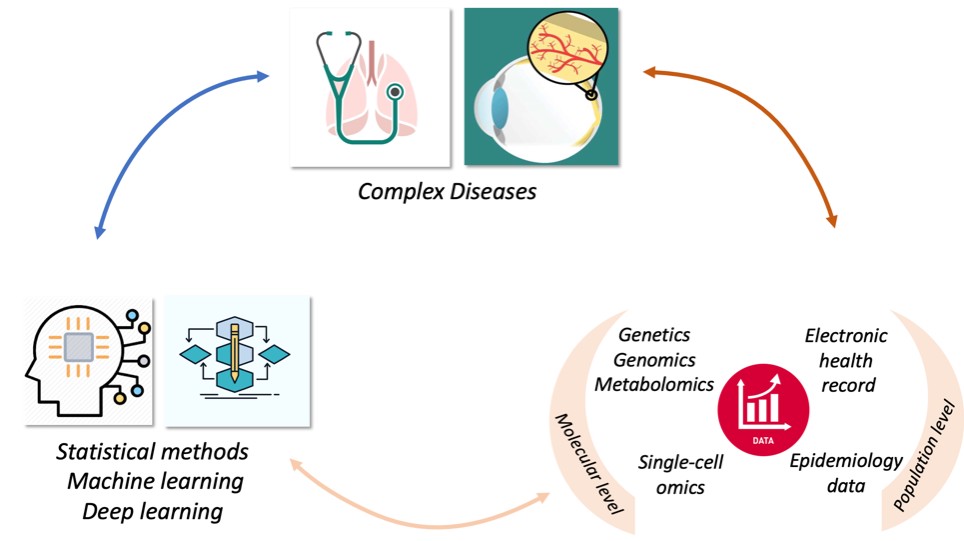
My lab focuses on developing statistical and computational methods for the analysis of large-scale biomedical data and applying them to study complex diseases. Collaborating with our colleagues, we use cutting-edge technologies including DNA-seq, RNA-seq, methylation, ATAC-seq and single cell sequencing to study human diseases such as childhood asthma and age-related macular degeneration. Chen lab welcomes collaborators and trainees to move science forward. Our lab is generously supported by National Institute of Health, National Science Foundation, University of Pittsburgh and UPMC.